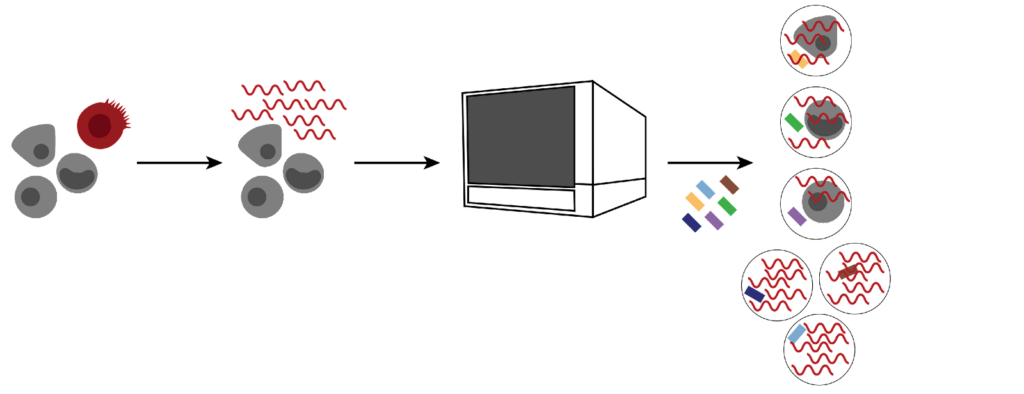
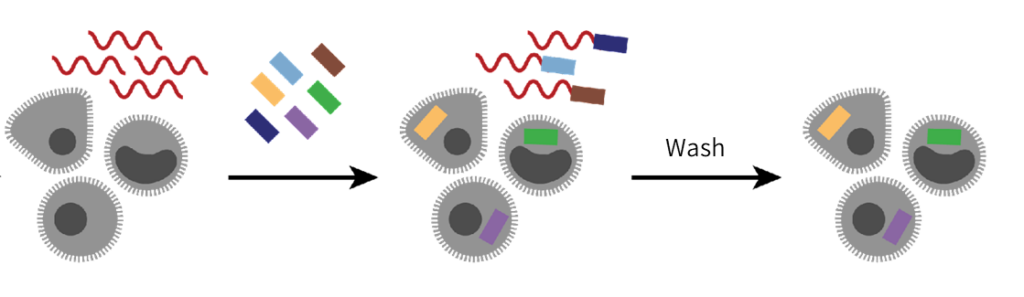
Sample source deserves specific consideration in sample preparation. Types of tissue cells, the amount and composition of the extracellular matrix, and the presence of necrotic cells will significantly impact the cell suspension protocol.
Because tissue cells interact via adhesion molecules and junctions, their disruption is necessary during dissociation or digestion. Considering the potential impact of sample preparation techniques on cell viability, RNA integrity, and gene expression, preparation protocols are often specific to the sample source.
Challenges exist in every sample, including some of the most used cell sources in scRNA-Seq experiments, such as:
Cell Lines
Certain cell lines grow adherent to the vessel. Gentle enzymatic treatments with TrypLE™ can effectively dissociate the cells from the culture surface without causing damage.
The density and confluence of cell cultures influence dissociation. High confluence or overcrowded cell cultures increase cell-cell adhesion, necessitating longer or more robust enzymatic reactions.
Some cell lines have resistant cell membranes, requiring prolonged incubation or more potent enzymatic treatments.
iPS Cell Colonies
Induced Pluripotent Stem cells (iPS) form densely packed colonies that clump together. When not adequately dissociated, these colonies form cell aggregates, which hinder efficient preparation.
Like other pluripotent stem cells, iPS cells maintain their pluripotency through interactions mediated by adhesion molecules. Researchers must employ enzymes or reagents specifically designed to target these adhesion molecules for successful dissociation.
Culture conditions and their differentiation states influence the sensitivity of iPS cells to cell preparation protocols. Under conditions that promote stronger cell-cell adhesion, such as when cultured in the presence of ROCK inhibitors, iPS cells may exhibit increased resistance to dissociation.
Brain Cells
Brain cells, particularly neurons, are large cells that exhibit intricate and highly specialized structures. Neurons’ extensive dendritic arborizations and long axons make the tissue particularly hard to handle.
A dense extracellular matrix (ECM) maintains brain tissue integrity by providing structural support. Additionally, cells are tightly connected through various adhesion molecules. While these structures are part of the integrity and function of neural circuits, they can impede digestion.
Neurons’ large size and the sternness of their scaffolding are common technical reasons for choosing to analyze single nuclei instead of whole cells, especially when the technology of choice is droplet-based microfluidics.
Indeed, the whole neuron’s size can be well above the droplet’s. Moreover, cell aggregates or the ECM components can clog the microfluidic channels affecting data quality.
Another challenge for brain cells is the presence of a myelin sheath. These lipid-rich membranes wrap around neurons multiple times, providing insulation and regulating cellular functions. These membranes can resist enzymatic digestion.
These membranes are naturally sticky, leading to obstruction when using droplet-based technologies, as they can adhere to the channels. Indeed, many protocols recommend incorporating a myelin removal step when working with brain cells.
Combinatorial barcoding is an instrument-free technology. Without the limitations of partitioning droplets and channels, there are no neuron size limits or risk of experimental failure due to obstructed channels.
Tumor Cells
While cell adhesion poses challenges in all tissues, it’s particularly problematic in tumor cells. The genetic and molecular alterations in tumor cells contribute to their cohesive and invasive properties. These alterations affect the expression of adhesion molecules and ECM components, making dissociation more difficult.
Tumor cells often exhibit fibrous or calcified regions, necrotic areas, or regions with high cellular density. Additionally, variations in lipid compositions and protein expressions impact membrane rigidity.
These characteristics collectively hinder efficient digestion and can result in incomplete dissociation of tumor cells.
Companies that sell tools and reagents for dissociation often offer tested protocols specifically designed for the challenging task of digesting tumor tissues.
Organoids
Organoids are three-dimensional structures that replicate the organization and functionality of corresponding in vivo tissues. They consist of diverse cell types embedded in an extracellular matrix of proteins and polysaccharides. Various cell-cell junctions interconnect these cells, providing structural support and facilitating communication.
Since organoids resemble organs in their cellular organization and structure, they encounter similar challenges during dissociation. Additionally, organoids consist of different cell types with varying sensitivities to dissociation methods.
Due to their complex nature, dissociation requires careful optimization and balance of enzymatic and mechanical conditions to preserve cell viability and cellular identity.