In the rapidly evolving world of medicine, the terms “antibody therapy” and “cellular therapy” have become synonymous with hope and promise.
Immunotherapies are a class of therapeutic treatments that enhance or suppress the immune system to fight severe diseases like cancer or autoimmune diseases respectively.
To accelerate and optimize immunotherapies, researchers increasingly use single-cell RNA sequencing (scRNA-seq) to examine the transcriptomics of individual immune cells. This technique identifies different cell types, including those that respond to a particular antigen, and uncovers both intercellular and intracellular communication, ultimately providing insights into the state of individual cells and the overall tissue environment.
This wealth of information — masked by bulk RNA sequencing — provides detailed insights into signaling pathways and molecular mechanisms. Concurrently characterizing B and T cell receptor (BCR and TCR) sequences allows for the investigation of dynamic changes in receptor-ligand interactions and the identification of clonal lineages that respond to specific antigens.
This comprehensive characterization is now considered essential for developing new therapeutic strategies. Such an endeavor requires scalability to achieve greater statistical power and capture full heterogeneity of the sample, and sensitivity to capture the nuances of expression changes and ease of implementation.
Traditional technologies lacking the necessary throughput, sensitivity, and cost-effectiveness, often fall short.
The adoption of innovative technologies that seamlessly integrate phenotypic and repertoire analysis at scale is crucial to propel the next wave of discoveries in immune-mediated diseases and enable the development of personalized therapies.
Global Growth of Immunotherapy-Based Clinical Trials
Antibody therapies are the fastest-growing class of drugs in oncology, autoimmunity, and chronic inflammatory diseases. The Cancer Research Institute (CRI), a nonprofit organization dedicated to the discovery and development of cancer immunotherapies, tracks the current Immuno-Oncology (IO) agents in clinical development and the clinical trials testing these drugs. The Global Immuno-Oncology Drug Development Pipeline shows a sharp increase in therapies between 2017 and 2020, with a 233% increase in just three years (fig. 1).
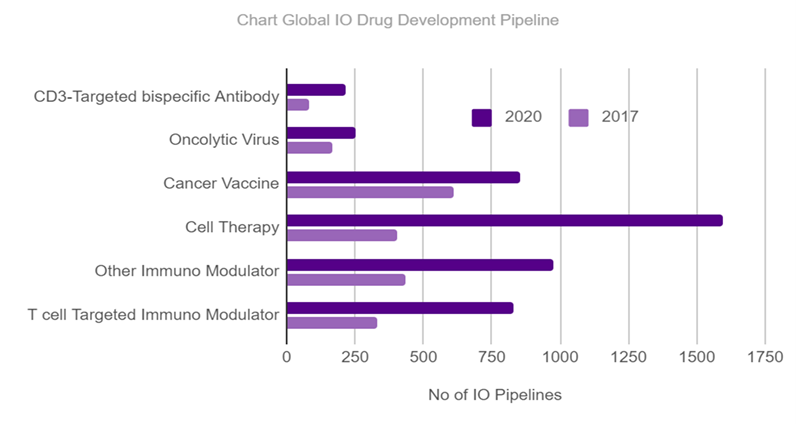
Fig. 1: A comparison of Immuno Oncology drug pipelines worldwide between 2017 and 2020 show a fast and exponential development of the number of immunotherapeutics. Data obtained from the CRI Anna- Maria Kellen Clinical Accelerator Team at CRI.
Moreover, in February 2022, the CRI updated the number of clinical trials started each year, reporting 5,683 trials worldwide, a 278% growth from 2017. While most of these trials included FDA-approved PD1/PDL1, a third of active trials tested other immunotherapy agents, with about 300 targets and pathways being investigated at the time of the report.
Immunotherapy has become a dynamic and rapidly advancing field in medicine, with intense competition to rapidly develop new therapies that harness the immune system to fight diseases.
Harnessing the Power of the Lymphocytes
The development of new immunotherapies greatly benefits from the ability to concurrently analyze the cellular transcription profile and the receptors’ paired sequences. Such capability empowers researchers to understand cell function, identify receptor variants with therapeutic potential, or match receptors with antigens to understand how immune cells recognize and respond to diseased cells.
Scrutinizing the Immune Response
Lymphocytes are highly heterogeneous, exhibiting a wide range of cell types and activation states. Traditional bulk sequencing methods mask this heterogeneity and provide an averaged view of the immune landscape.
In contrast, single cell sequencing allows for the identification and characterization of distinct lymphocytes subpopulations, each with its unique functional properties and roles in the immune response.
Additionally, the genes encoding the paired chains that constitute the TCR and BCR repertoires are distributed across different chromosomes reflecting their physical separation. Prior to the advent of scRNA-seq, high throughput bulk sequencing methods aggregated data from millions of cells, concealing the pairing structure of the chains.
Single cell resolution enables the sequencing of both chains of the same receptor.
Paired TCR and BCR receptors chain analysis reveals the full diversity of the immune repertoire, the dynamics between immune cell networks and their interactions with surrounding tissues, revealing how lymphocytes recognize and engage specific ligands.
This comprehensive view of the immune system’s intricate dynamics and its targeted response to antigens is key to discovering new targets for cell and antibody therapies.
Quantify and Track Clonotypes
One of the significant challenges in cell therapies, particularly in the context of adoptive cell therapies like enhanced TCR-T and CAR-T cell therapies, is the short-term persistence of disease-reactive clones in the host after infusion.
The ability to sequence paired chains enables researchers to discover, detect, and track the clonal expansion of immune cells, and the stability of infused engineered T cells over long periods of time.
For example, scRNA-seq in long term studies have helped demonstrate the clonal stability of infused CAR-T cells, with ongoing functional activation and proliferation, providing insight into the CAR T cell characteristics associated with anti-cancer response and long-term remission in leukemia. Tracking engineered T cell clones can help optimize cellular doses and predict the response.
Moreover, the information gained by targeting specific immune cell populations can be used to develop new therapeutics, such as vaccines or immunotherapies, or even resistance to such therapies. By exploring the antigen expression patterns in healthy and malignant cells, researchers have uncovered factors that drive resistance to engineered T cell therapies in multiple myeloma.
Effective long term research strategies should incorporate advanced technologies designed to fix and preserve the transcriptional integrity of precious patients’-derived cells. This approach is crucial to obtain a comprehensive profiling of the clonal diversity.
Large Scale Antibody Discovery
Antibodies are critical tools in research, diagnostic, and therapeutic fields.
The success of existing antibody therapies has triggered the demand for more. Their discovery and use has been further propelled by the need for a rapid response to the emergence of new diseases.
When scRNA-seq is run concurrently with repertoire sequencing, it can recover a larger number of antibody lineages compared to previous technologies.
In this paper, researchers have leveraged scRNA-seq and BCR sequencing to discover hundreds of potent neutralizing antibodies targeting SARS-CoV-2 Omicron variants. By analyzing over 277,000 B cells, they identified a set of memory B cell-derived antibody candidates. Such findings pave the way for further research into their binding and neutralizing capabilities against various SARS-CoV-2 variants, potentially offering new avenues for COVID-19 treatment and prevention.
This study utilizes extensive resources to process a quarter of a million cells, underscoring the necessity for more resource-efficient methodologies.
Indeed, conducting such experiments using traditional scRNA-seq technologies challenges the concept of scalability and ease of implementation. While these large repertoire studies are better positioned to provide insights into therapeutic interventions and biomarkers, they also face higher costs that can make funding difficult to obtain. This could potentially discourage scientists from initiating extensive studies.
Fortunately, today, new cutting edge technologies such as combinatorial barcoding are streamlining high throughput scRNA-seq and immune profiling experiments by profiling up to 1 million cells in one experiment, with remarkable efficiency and sensitivity, and significantly reducing the logistical footprint of such experiments.
The Parse Biosciences Evercode Assay: A Game-Changer
As immune-based treatments represent one of the fastest-growing classes of newly FDA-approved therapeutics worldwide, researchers who aspire to cross the finish line in the race for the next successful clinical trial need high-throughput, sensitive, easy to implement, and scalable scRNA-seq technologies.
The Parse Biosciences’ Evercode kits have all these advantages built into the workflow.
Unlike droplet-based methods, the Evercode fixation-based, split-pool combinatorial barcoding technology can dramatically scale up scRNA-seq experiments without the need for dedicated instruments. With only a thermal cycler, a centrifuge, and some pipettes, users can sequence up to 1 million cells or nuclei in 96 samples. Such a dramatic increase of cells and samples throughput reduces time and resource barriers.
The technology is fixation-based, allowing the accrual and storage of samples until they are ready to be processed together at a convenient time. This capability enables researchers to discover and track rare clones and cell states over extended periods with unprecedented resolution and no concerns for batch effects.
Recently, the Evercode assay has been expanded to capture BCR and TCR sequences along with whole transcriptome expression from the same cells.
This groundbreaking method reveals the complexity of the immune repertoire by detecting over 90% of the paired chains in both the TCR and the BCR (Fig. 2).
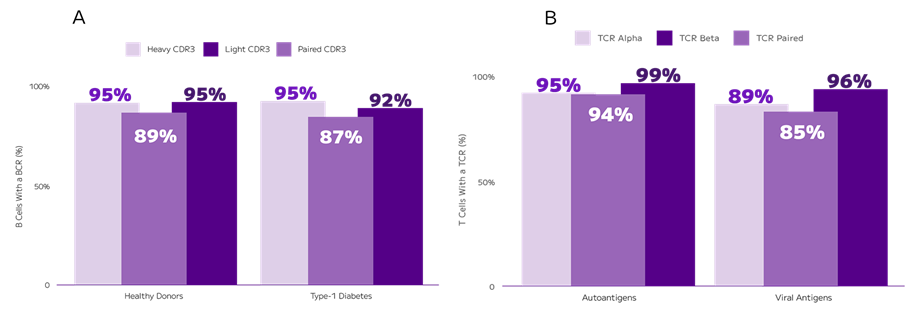
Fig. 2: Understanding the immune repertoire diversity necessitates sensitive detection of TCRs and BCRs along with whole transcriptome profiles. In an experiment comparing T1D and control donors, Evercode BCR captured both heavy and light chains with 87-89% BCR pairing efficiency (A). In an experiment profiling viral or auto antigen stimulated T cells, Evercode TCR captured paired alpha and beta chains in 85-94% of cells (B).
Evercode BCR and Evercode TCR in Action
Scientists in major universities are already using the Evercode BCR Evercode TCR assays.
The Boyd Lab at Stanford University used the Evercode BCR assay to capture both the cellular makeup and BCR repertoire in healthy people. They fixed the B cells and later ran the assay. The whole transcriptome data showed high transcripts and gene detection across 87,000 cells. The paired heavy and light chains generated across 82,000 B cells featured stimulated B cells that differentiated and underwent isotype switching, resulting in both memory B cells and antibody secreting plasma cells (Fig. 4).
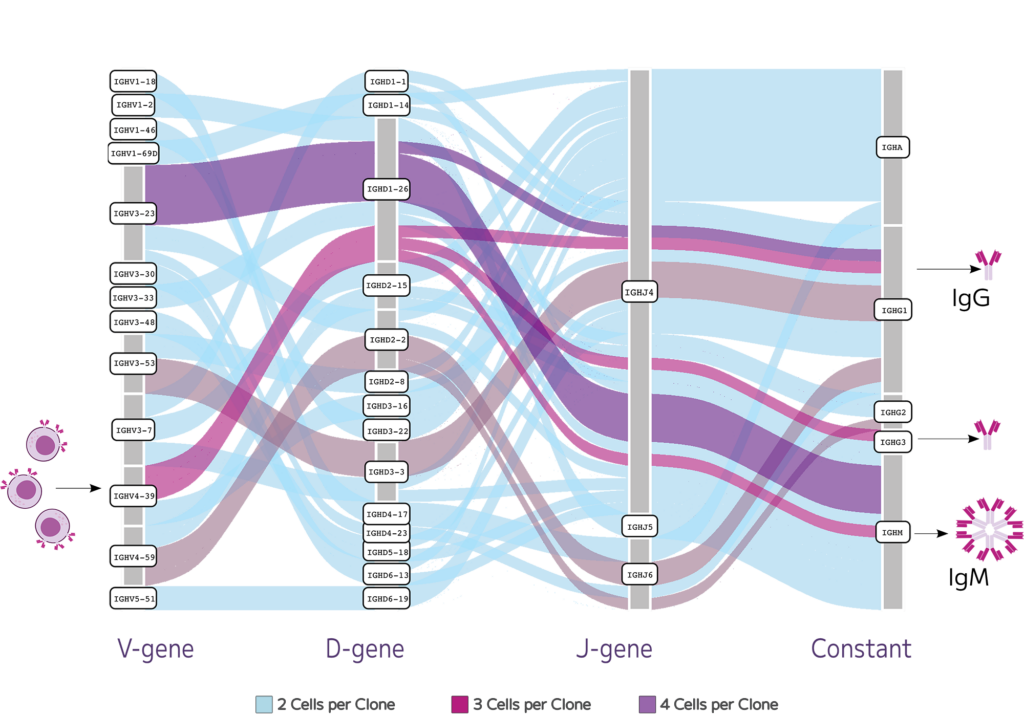
Fig.4: Stimulated B cells can differentiate and undergo isotype switching resulting in both memory B cells and antibody secreting plasma cells. Evercode Fixation preserves fragile cell types like plasma cells, enabling deep profiling of the clonal diversity and providing insight into patient specific immune responses.
The Reticker-Flynn Lab at Stanford is utilizing the Evercode TCR technology to examine the impact of the lymph nodes in tumor metastasis and subsequent changes to the phenotype and clonotype diversity within mouse T cells. The study is ongoing, and the team plans to leverage TCR sequencing data to gain insights into tumor dynamics and activity.
Moreover, in a recent study presented at AAI 2024, Parse scientists demonstrated the Evercode power of scalability by examining BCR clonal diversity in autoimmune diseases using the Evercode BCR assay. They simultaneously profiled both the BCR and whole transcriptome of up to 1 million cells from 24 samples in a single experiment. This approach enabled the detection of all major B cell subtypes in the peripheral blood of 12 subjects with autoimmune diseases and 12 healthy controls. The data illustrated BCR isotypes, captured a majority of cells with a paired chain, detected full length chains, and differentiated dominant and rare clones across a host of autoimmune diseases (Fig. 5).
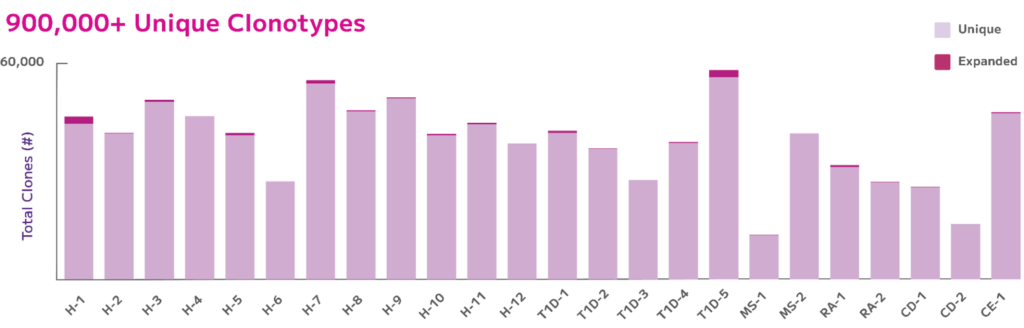
Fig. 5: Number of Unique vs. Expanded Clones. Total unique clones varied between 13,000 – 59,000 per donor where all donors had over 95% unique clones in their clone pool with donor H-1 and T1D-5 having the most expanded clones ~5%.
When focusing on the 5 samples from Type 1 Diabetes donors, the team recognized different immune responses in these patients based on their clonal expansion and isotype switch.
At the same conference, Parse scientists demonstrated the sensitivity, accuracy, and scalability of the Evercode TCR analyzing the effects of aging on mouse T cell receptor diversity using the Evercode TCR assay. By profiling hundreds of thousands of mouse spleen-resident T cells at different life stages, they identified all major T cell populations and their paired TCR sequences. These findings revealed differences in clonotype diversity correlated with age and cell type, highlighting the utility of this technology in studying the impact of aging on the immune system (Fig. 6).
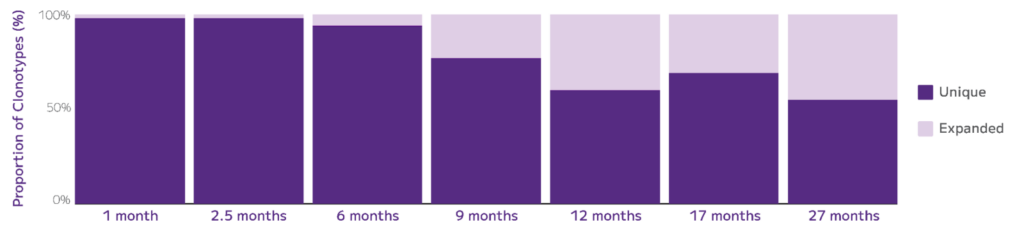
Fig. 6: Evercode TCR for Mouse: proportion of unique to expanded clonotypes over the mouse lifespan. The unique clonotypes (dark purple) are defined as only being detected in 1 cell: their percentages decrease from 98% to 55% over 26 months, demonstrating the T cell repertoire contraction over time.
Conclusion
The advancements in immunotherapies and the corresponding increase in clinical trials reflect the significant progress in this field. The upward trend in market growth from 2023 to 2031 aligns with the increasing demand for personalized medicine and the integration of advanced technologies in research.
Scientists who want to contribute effectively to these demands need access to scalable technologies that can produce large datasets capable of capturing the fine aspects of the immune repertoires.
Fortunately, Parse Biosciences has developed Evercode, which can be seamlessly integrated into any laboratory to accommodate the pressing demands for large scale immune profiling projects.
The Evercode BCR and the Evercode TCR pair with the urgency of the scientific and medical community to drive forward the discovery and development of targeted and personalized therapies.