Performance of Immune1000 Gene Select in Human Immune Cells (PBMCs)
Key takeaways
- High resolution and cell type proportion after enrichment
- >10 times less sequencing required
- High on-target rate (>80%) with Gene Select
High resolution data with less sequencing
Evercode WT libraries and the same libraries enriched with Immune1000 Gene Select showed similar clustering resolution, and cell type proportion with >10 times less sequencing depth. The Immune1000 Panel contains probes designed to enrich for immune cell type markers and pathways.
Immune1000 Panel was used to enrich 8 sublibraries from an Evercode WT kit. Libraries were then sequenced at 2,000 reads/cell, yielding an on-target rate of >80%. When compared to the non-enriched libraries sequenced at 28,000 reads/cell, the Immune1000 panel showed the same cell type clustering as the whole transcriptome.
Figure 1. Cell type clustering in non-enriched and enriched libraries. Whole transcriptome libraries were prepared from human peripheral blood mononuclear cells (PBMC) obtained from 2 donors. Immune1000 captured immune specific genes from 105,462 cells across 8 sublibraries.
Concordant cell types
A correlation plot produced unbiased concordance of immune cell types from 2 donors in libraries prepared with Evercode WT and enriched with Immune1000 Gene Capture (r2=0.998).
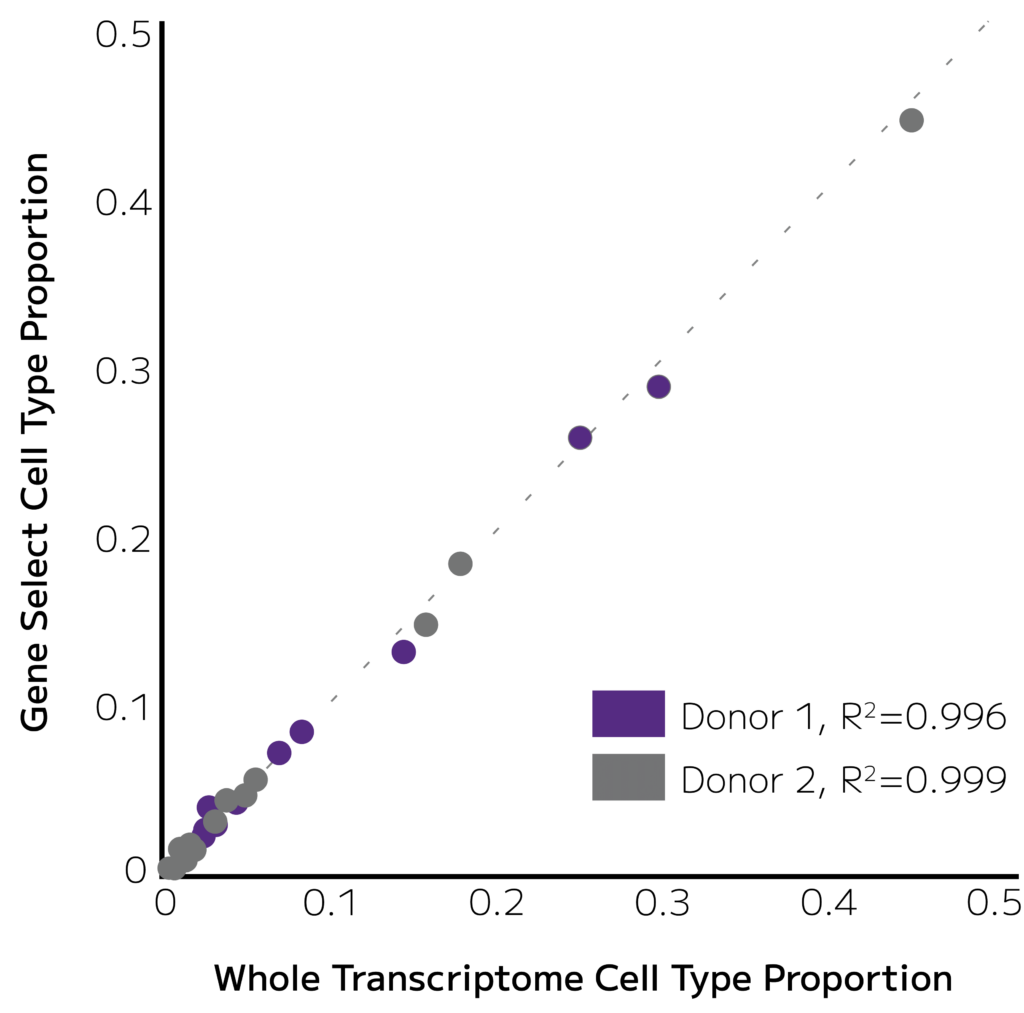
Figure 2. Cell Type proportions are retained. Evercode WT libraries with and without Immune1000 Gene Capture show a nearly perfect concordance of cell types. Each dot corresponds to an immune cell type.
Next Steps
Download the data files generated from the preparation and analysis of two donors’ PBMC samples. The datasets include the experimental summary reports, the digital gene expression matrix, the gene files, and the cell metadata with and without Immune1000 Gene Capture.
Downloads
We're your partners in single cell
Reach out for a quote or for help planning your next experiment.