Performance of Evercode™ WT in Human Immune Cells (PBMCs)
Key takeaways
- Sensitivity increased in PBMC samples
- More consistent gene detection
- Unbiased gene expression
More genes with less sequencing
Evercode WT v2 detected substantially more genes and transcripts in human peripheral blood mononuclear cells (PBMC) compared to Evercode WT v1. To compare, four PBMC samples were fixed and prepared in parallel with Evercode WT v1 and Evercode WT v2.
A significant increase in genes was detected with Evercode WT v2 compared to Evercode WT v1. The increased gene detection as plotted against the sequencing depth is shown below (Figure 1). At 100k reads/cell, Evercode WT v2 had 63% more genes detected. At 20k reads/cell, 48% more genes were detected than Evercode WT v1.
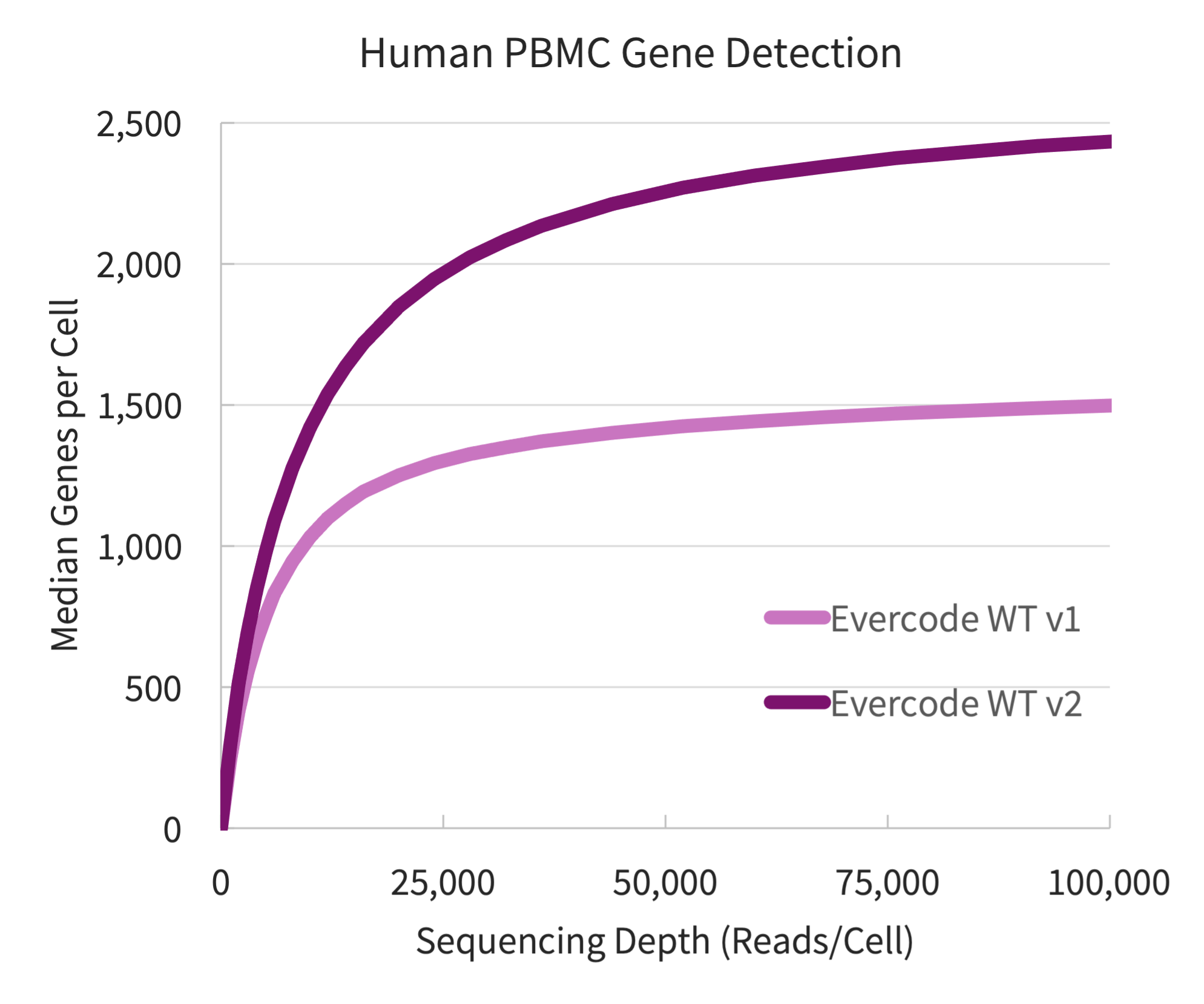
Figure 1. Gene Detection for Human PBMCs. Gene detection from four fixed PBMC samples prepared using Evercode WT v1 and Evercode WT v2, in parallel. A sublibrary from each experiment was sequenced and processed using the Parse Biosciences data analysis pipeline.
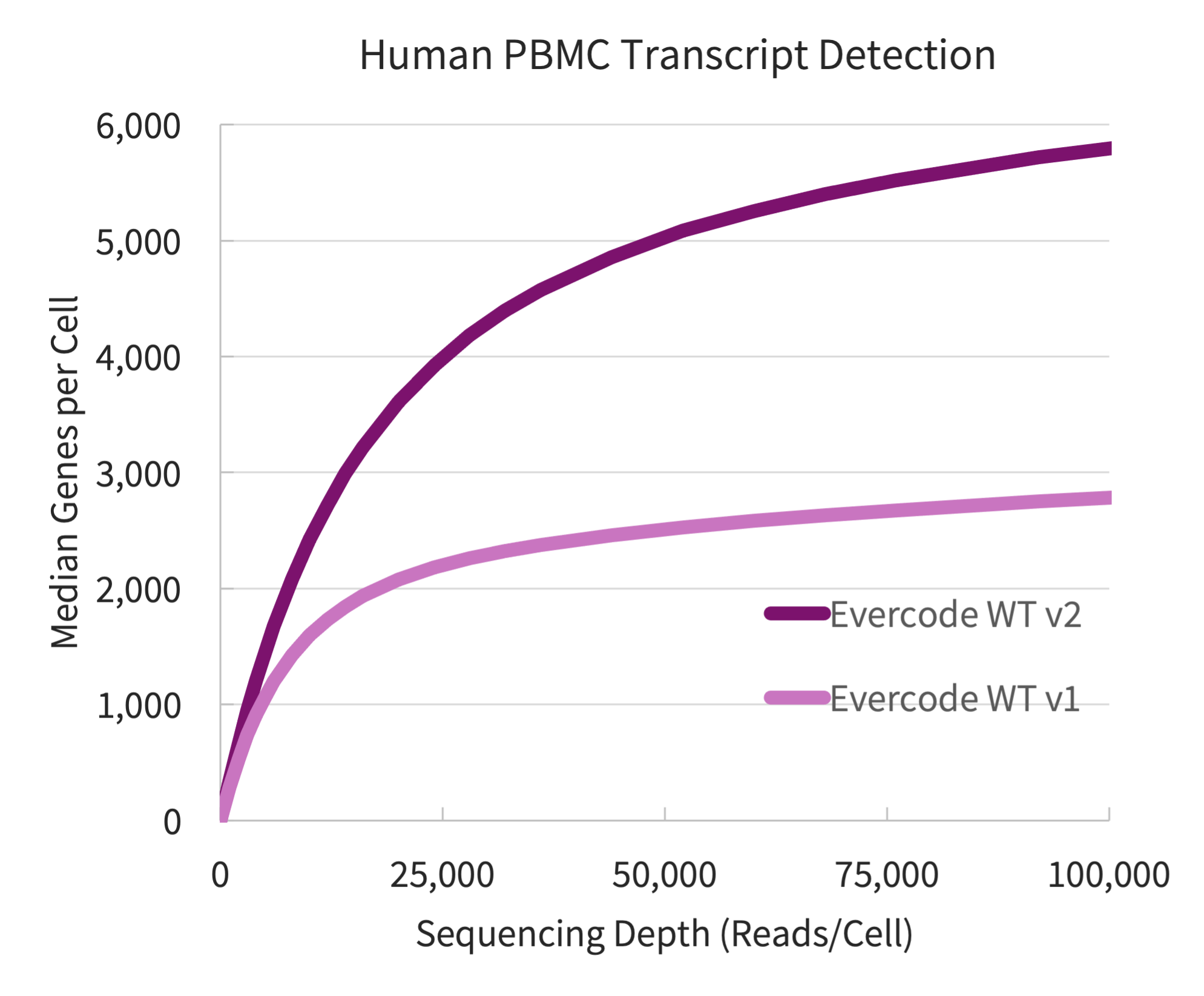
Figure 2. Transcript Detection for Human PBMCs. Transcript detection from four fixed PBMC samples prepared using Evercode WT v1 and Evercode WT v2, in parallel. A sublibrary from each experiment was sequenced and processed using the Parse Biosciences data analysis pipeline.
More consistent gene detection
Consistent gene detection between samples results in fewer sample failures, more predictable sequencing planning , and straightforward data analysis. The reproducibility and distribution of the genes per cell for Evercode WT v1 and Evercode WT v2 were plotted (Figure 3). The medians were more consistent across samples prepared with Evercode WT v2.
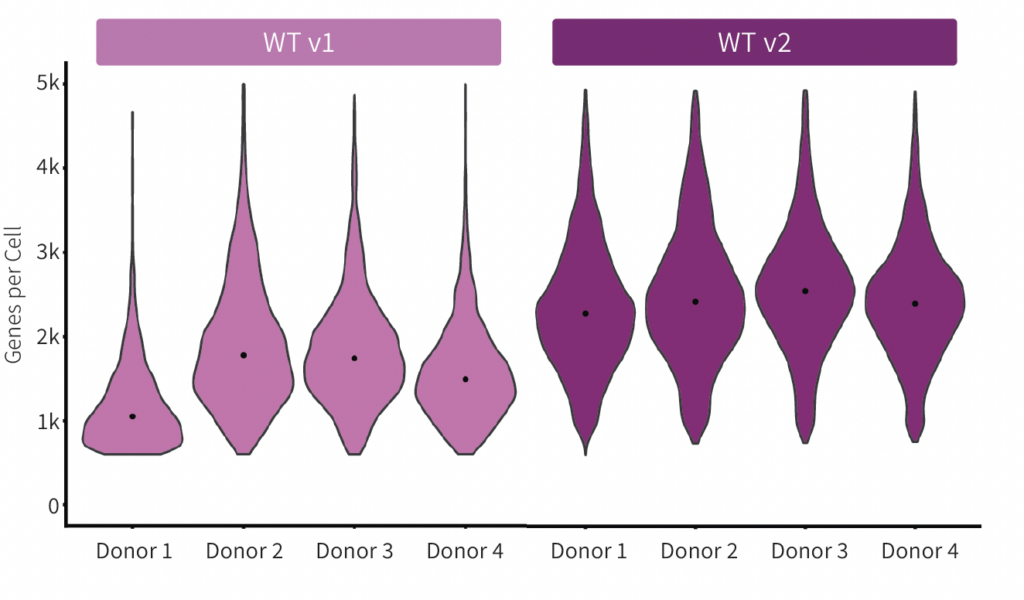
Figure 3. Gene Detection Across Samples for Human PBMCs. Violin plot showing the number of detected genes per donor for Evercode WT v1 (left) and Evercode WT v2 (right). The dot (black) denotes the median genes detected per donor.
Unbiased gene expression
Gene correlation plots confirm unbiased gene expression of the PBMC samples prepared with Evercode WT v1 and Evercode WT v2 (r2 = 0.982) (Figure 4).
Researchers can transition to Evercode Whole Transcriptome Solution v2 kits to benefit from greater gene sensitivity and improved sample robustness while integrating previous data into the ongoing studies.
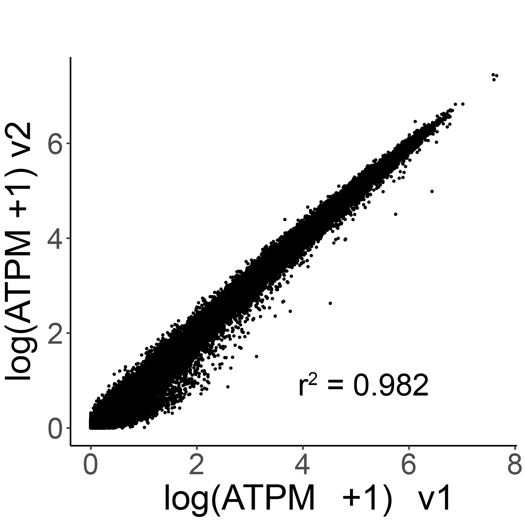
Figure 4. Gene Expression Correlation Between Evercode WT v1 and Evercode WT v2 in Human PBMCs. The average gene expression (log average transcripts per million) between the Evercode WT v2 and Evercode WT v1 results was compared. The r 2value indicates a high degree of correlation in gene expression between Evercode WT chemistry versions.
Next Steps
- Download the data files generated from the preparation and analysis of the adult mouse liver nuclei samples. These datasets include the experimental summary report, the digital gene matrix file, the cell metadata file, and genes file.
- Explore more by reviewing the full experimental plan and detailed results across multiple sample types in the bioRxiv paper.
We're your partners in single cell
Reach out for a quote or for help planning your next experiment.