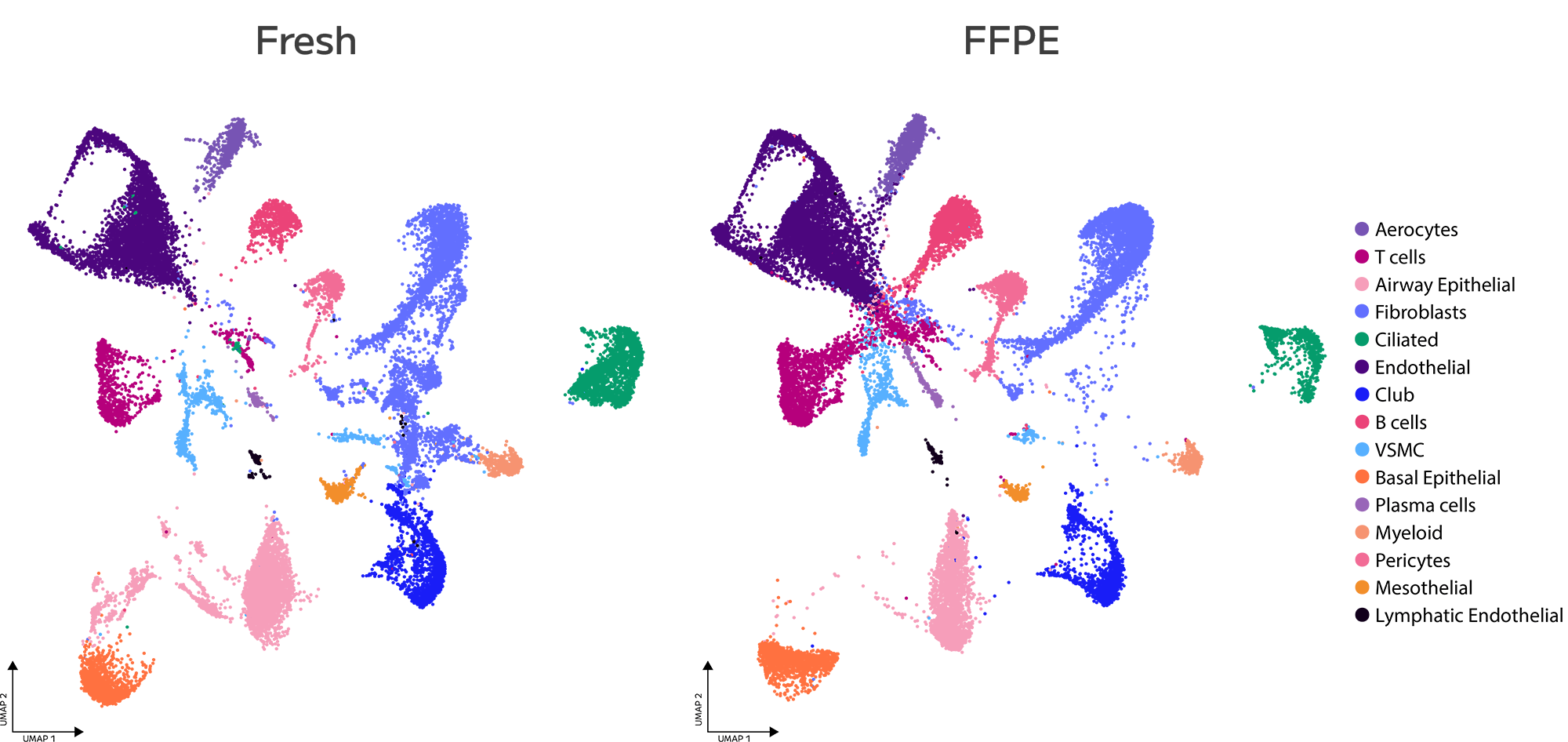Technology 
Accessible single cell
The end-to-end solution
Our solution takes you from single cell or single-nuclei suspension through library prep and sequencing and delivers immediate results via our analysis software, Trailmaker.
製品 
Now Available
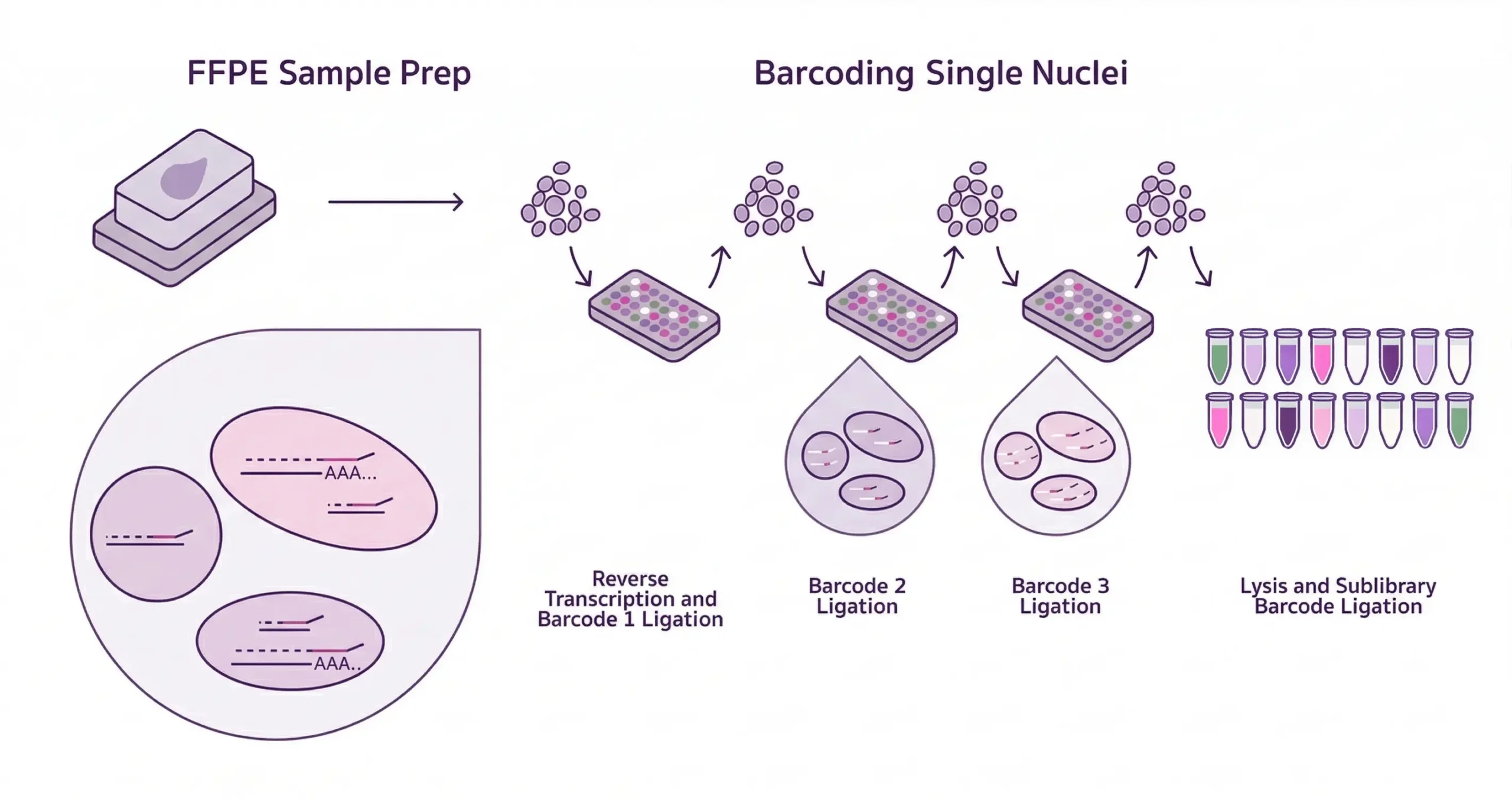
Evercode™ WT FFPE
With Evercode WT FFPE, researchers can see the complete molecular landscape of each cell without the constraints of predefined probe panels.
会社情報